- New study used Federated Learning to predict severity of Covid-19 for E.R. patients.
- Significant improvements seen in central vs. local models.
- Model slated for use in production in the near future.
Many ethical and legal challenges surround COVID-19 data analysis, including data ownership, data security, and privacy issues. As a result, healthcare providers have typically preferred models validated on their own data. However, this limits the scope of analysis that can be performed, often resulting in AI models that lack diversity, suffer from overfitting, and demonstrate poor generalization. One recent study titled Federated learning for predicting clinical outcomes in patients with COVID-19, published in September 15 issue of Nature Medicine [1], offered a solution to these problems: Federated Learning (FL).
FL is a privacy-protection model trained in heterogeneous, distributed networks [2]. It allows model training from diverse sources without data centralization; The AI models are trained from disparate sources, while the data itself remains in the original location.
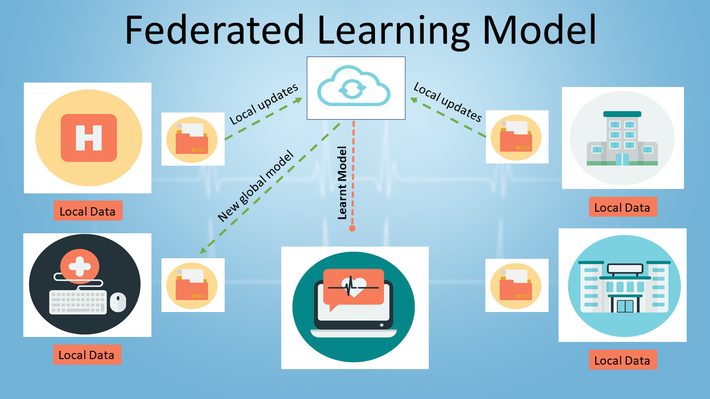
Client-Server FL Model in Healthcare Settings
In FL models, hospitals and other healthcare settings can be thought of as remote devices, containing a wealth of patient data. This patient data is usually required to remain local due to privacy concerns and various administrative, ethical, and legal constraints. The study authors used a particular FL approach called client-server, where clients download a trainable model from a centralized (federated) server. The model is updated on the remote devices with new data and partial training tasks, The updated model is then uploaded back to the central server, where multiple client updates are merged, thus improving the model. The process continues iteratively until training is complete.
Privacy concerns were mostly alleviated during the study because the data remained on the local servers; The communication between the central serves and local nodes contained only model weights or gradients. The authors added an additional layer of security to avoid the risk of model inversion or reconstruction of training images.
The Study
The study goal was to enable healthcare facilities to triage patients more effectively. The research involved a range of data gathered from multiple sites across four continents. In total, 20 features were input to the model, including patient age, lab results, and imaging. Outcomes were measured based on the type of oxygen therapy a patient required in a predefined prediction window24 to 72 hours from when the patient arrived in the E.R. For example, mild cases of Covid-19 did not need any supplemental oxygen, while more severe cases required mechanical ventilation. The result was a risk score, from 0 to 1, which predicted patient outcomes; A risk score of zero indicated no supplemental oxygen, while deaths were recorded as a risk score of 1.
The hope was that that the FL model would perform better than local models, giving better generalization across healthcare systems.
Results
As well as being one of the first FL models for COVID-19, the risk model was also one of the largesttraining on data from 16,148 cases of Covid-19 hospital admissions. The model was compared with locally trained models on the clients servers. Validation was performed at various hospitals in Massachusetts.
The authors recorded significant improvements over locally trained models, including a 38% improvement of generalizability. The FL model had an increased ability to predict mechanical ventilation or death with a sensitivity (the ability to detect true positives) ranging from .95 to 1. The following image, from the study, shows AUC scores for the local vs. central models:
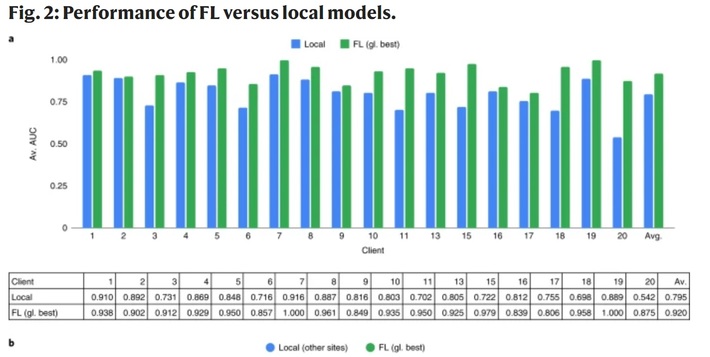
The conclusion from the study was that the FL model captured more diversity than locally trained models, setting the stage for real-world use in ERs for evaluating risks of serious Covid-19 outcomes in ER patients. Further studies are underway to find optimal training parameters for individual client sites; These may incorporate automated hyperparameter searching, neural architecture search, and other ML approaches.
References
FL: Image by Author (Icons: Adobe Creative Cloud).
[1] Federated learning for predicting clinical outcomes in patients wit…
[2] https://blog.ml.cmu.edu/2019/11/12/federated-learning-challenges-me…
